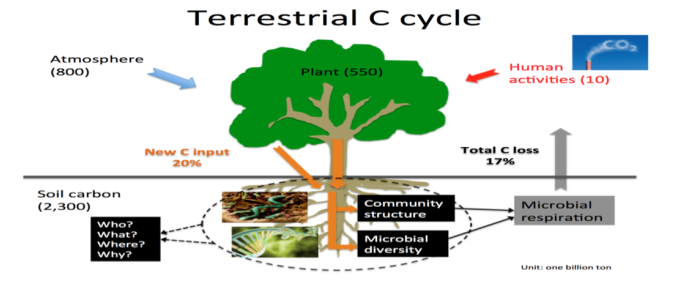
Microbial communities modulate nutrient transformation and storage in ecosystems. However, biogeochemical approaches (e.g., stable isotope probing) always focus on process, pools, and fluxes, whereas molecular techniques (genomics, transcriptomics, proteomics, metabolomics) usually focus on individual or community diversity, composition, and community structure. I am trying to combine the two approaches to improve our understanding of microbial processes (shifts, turnover, evolution, etc.) that control biogeochemical cycling in soils. My research centers on microbial community responses and microbial-mediated processes controlling carbon and nutrient cycling feedbacks of biosphere, and their effects on ecosystem functions, especially under the increasing climate change (i.e., warming, drought, etc). These techniques can also be applied to agricultural ecosystems, especially in the plant-soil interactions, to improve crop production, increase nutrients, and reduce potential environmental pollution.
Major research interests:
1. How are biogeochemical C and N cycling affected by
2. What are the relationships among substrate inputs (energy, nutrients), soil C or N cycling, and long-term C and N storage?
3. How can we better model and predict nutrient and C cycling in a changing world?
4. What are the plant-soil relationships?
Major research interests:
1. How are biogeochemical C and N cycling affected by
- Environmental factors: temperature, precipitation.
- Soil biological factors: different groups of microorganisms like bacteria, fungi, etc.
- Soil physical factors: soil structure, texture, such as clay minerals, aggregates, etc.
- Soil chemical factors: different complexity of soil organic matter (SOM), such as humic substances, easily decomposable sugars or other components.
- Plant factors: plant litter inputs, rhizosphere root exudates, plant-microbe-soil interactions, etc.
2. What are the relationships among substrate inputs (energy, nutrients), soil C or N cycling, and long-term C and N storage?
- The relationship could be modulated by substrate types, amounts, addition frequency, and time periods.
- Soil enzyme functions, especially those genes encoding C and N metabolic activities.
- Biodiversity of microbial taxa and functional traits.
- Techniques such as isotopes (13C, 18O, 15N, 2H), multi-omics, and NanoSIMS can be used to verify the relationships between substrate inputs and C or nutrient cycling.
3. How can we better model and predict nutrient and C cycling in a changing world?
- Microbe-based models, biogeochemistry-based models, Ecosystem models.
- Add microbial traits, omics data, enzyme activity to different models to select the best ones that can more accurately predict nutrient and C cycling.
4. What are the plant-soil relationships?
- With environmental stressors, how do plants and soil microbes respond?
- Could we select drought-resistant plant or soil communities to increase plant fitness or crop yields? Or increase plant access to more nutrients, or reduce pathogens that appear with climate change?
- Climate change likely will affect soil health indicators, such as nutrient cycling, water use, soil physical structure and chemistry, and diversity.